1 Data to Analyze
To answer the research question “Does obesity increase the risk of developing diabetes?” in the U.S. context, we do the following:
1.1 Choose a U.S. data source
- Data source: National Health and Nutrition Examination Survey (NHANES) (Disease Control and Prevention 2021)
- Availability: NHANES is a publicly available dataset that can be downloaded for free from the CDC website.
- Design: Observational cross-sectional data. Hence, inferring causality is not a possibility or our objective here.
1.2 Confounder identification
Directed acyclic graph (DAG)
flowchart TB A[Obesity A] --> Y(Diabetes Y) L[Confounders C] --> Y L --> A
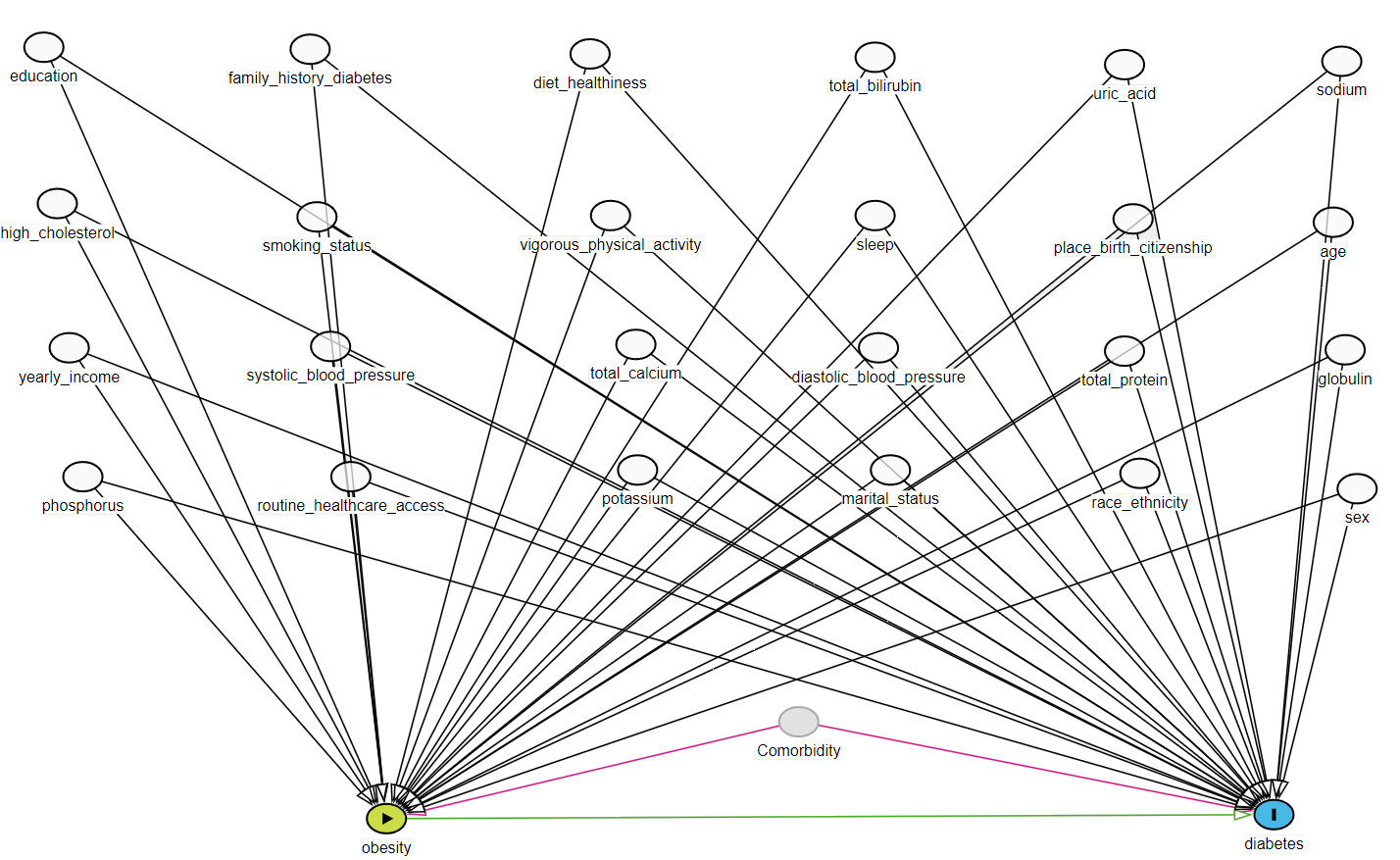
In standard Propensity Score analysis, we rely on Investigator-Specified Covariates. According to causal theory, we should include:
- ✅ Confounders: Variables causing both Exposure and Outcome.
- ✅ Risk Factors: Variables causing the Outcome (to improve precision)
We must avoid:
- ❌ Instruments: Variables causing Exposure only (increases variance/bias).
- ❌ Colliders: Effects of both Exposure and Outcome (induces bias).
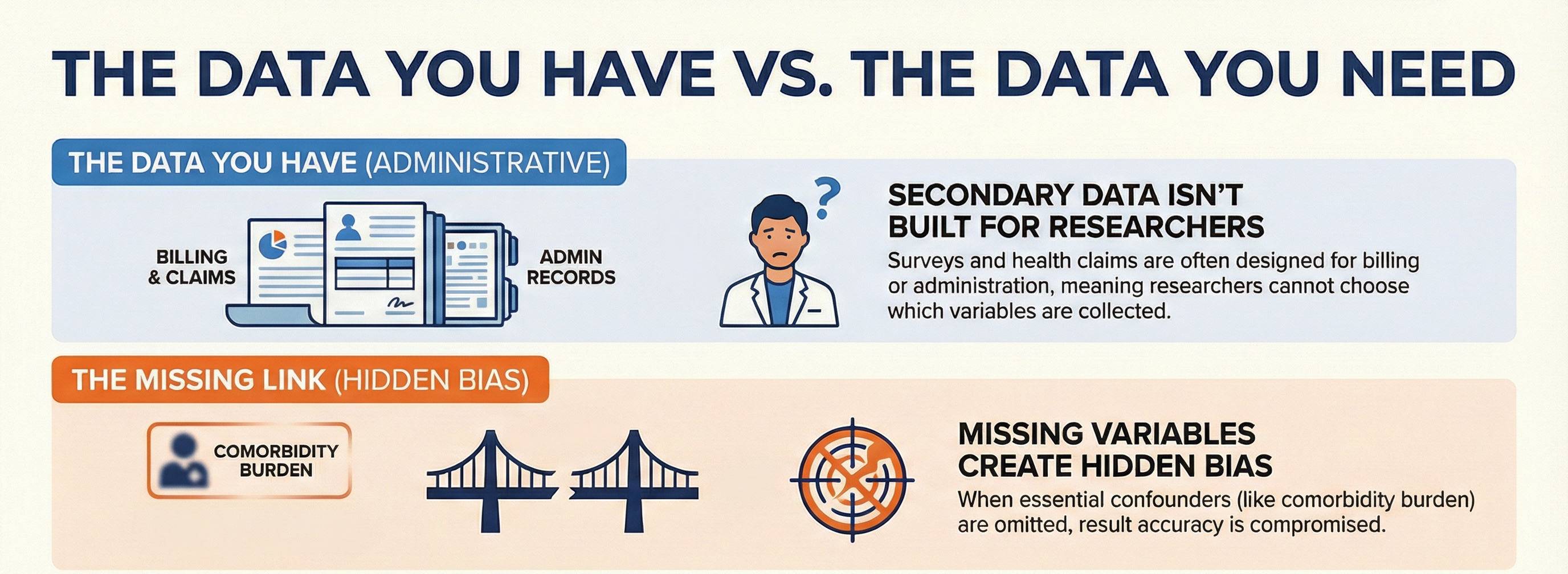
When using secondary data sources such as NHANES or health administrative databases, the data is often collected for administrative, billing or general monitoring rather than specific research questions. Because researchers do not have control over which variables are included in the survey/questionnaire, essential variables required to fully adjust for bias (confounders) may be missing.

Exposure: Being obese
Outcome: Developing diabetes
Confounders: Demographic and lab variables
1.3 Structure of the data
flowchart LR D[NHANES 2013-14] --> demo[Demographic Variables and Sample Weights] demo --> Age demo --> Sex demo --> Education demo --> r[Race or ethnicity] demo --> m[Marital status] demo --> Income demo --> b[Birth place] demo --> sf[Survey features: sampling weights, strata, cluster] D --> bmi[Body Measures] bmi --> Obesity D --> diq[Diabetes] diq --> Diabetes diq --> f[Family history of diabetes] D --> smq[Smoking - Cigarette Use] smq --> Smoking D --> dbq[Diet Behavior & Nutrition] dbq --> Diet D --> paq[Physical Activity] paq --> p[Physical activities] D --> huq[Hospital Utilization & Access to Care] huq --> mm[Medical access] D --> bpx[Blood Pressure] bpx --> sbp[Systolic Blood Pressure] bpx --> dbp[Diastolic Blood Pressure] D --> bpq[Blood Pressure & Cholesterol] bpq --> hc[High cholesterol] D --> slq[Sleep Disorders] slq --> Sleep D --> biopro[Standard Biochemistry Profile] biopro --> u[Uric acid] biopro --> Protein biopro --> Bilirubin biopro --> Phosphorus biopro --> Sodium biopro --> Potassium biopro --> Globulin biopro --> Calcium D --> rxq[Prescription Medications - ICD-10-CM codes] style D fill:#FFA500; style rxq fill:#00FF00; style biopro fill:#00FF00; style slq fill:#00FF00; style bpq fill:#00FF00; style bpx fill:#00FF00; style huq fill:#00FF00; style paq fill:#00FF00; style dbq fill:#00FF00; style smq fill:#00FF00; style diq fill:#00FF00; style bmi fill:#00FF00; style demo fill:#00FF00;
We do the same for the following cycles:
- NHANES 2015-16
- NHANES 2017-18
1.4 Identify measured and unmeasured variables in the data
Find variables capturing the following concepts in the data based on a hypothesized DAG.
| Role | Data Component | Variables considered based on DAG |
|---|---|---|
| Outcome | DIQ | Have diabetes1 |
| Exposure | BMX | Obese; BMI >= 30 |
| Confounder | (demographic) DEMO | Age, Sex, Education, Race/ethnicity, Marital status, Annual household income, County of birth, Survey cycle year |
| (behaviour) SMQ, PAQ, SLQ, DBQ | Smoking2, Vigorous work activity, Sleep3, Diet4 | |
| (health history / access) DIQ, HUQ | Diabetes family history, Access to care5 | |
| (lab) BPX, BPQ, BIOPRO | Blood pressure (systolic, diastolic6), Cholesterol, Uric acid, Total Protein, Total Bilirubin, Phosphorus, Sodium, Potassium, Globulin, Total Calcium |
- 14 demographic, behavioral, health history related variables
- Mostly categorical
- 11 lab variables
- Mostly continuous
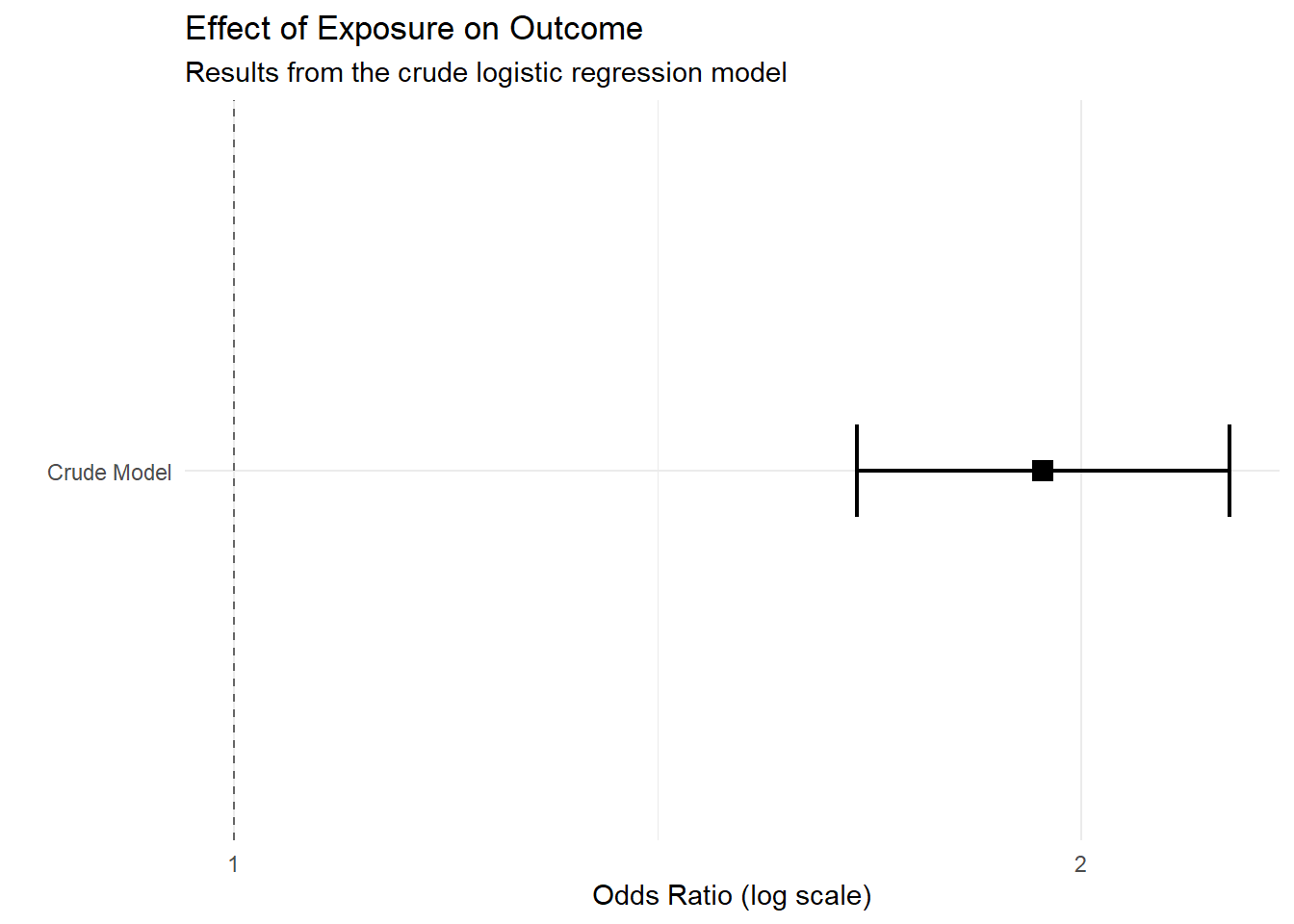
1.5 Fitting crude model to obtain OR
Here we estimate the crude association between the exposure and the outcome.
out.formula <- as.formula("outcome ~ exposure")
fit <- glm(out.formula,
data = hdps.data,
family= binomial(link = "logit"))
fit.summary <- summary(fit)$coef["exposure",
c("Estimate",
"Std. Error",
"Pr(>|z|)")]
fit.ci <- confint(fit, level = 0.95)["exposure", ]
fit.summary_with_ci.crude <- c(fit.summary, fit.ci)
knitr::kable(t(round(fit.summary_with_ci.crude, 2)))| Estimate | Std. Error | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|
| 0.66 | 0.08 | 0 | 0.51 | 0.81 |